- ID: DS-L02
- Type: Lesson
- Audience: Beginner / Intermediate
- Theme: Exploration before interpretation
Analytical work does not begin with modeling.
It begins with looking carefully at the data.
Before fitting models or drawing conclusions, we need to understand:
- what the dataset contains
- how variables are structured
- whether values look plausible
- whether early patterns deserve closer attention
In CDI, this stage is not a formality. It is where analytical judgment begins.
In this lesson, we work with the classic Iris dataset (Fisher 1936), which contains measurements of flower characteristics across three species.
You will:
- load the dataset into a pandas DataFrame
- inspect its structure and column types
- generate summary statistics
- create exploratory visualizations
- save the dataset for reuse in later lessons
Why this step matters
A dataset can appear clean at first glance and still contain issues that affect later analysis.
Early exploration helps answer practical questions:
- how many rows and columns are present?
- which variables are numeric and which are categorical?
- do any values look unusual?
- are there visible group differences?
- what should be examined more carefully next?
This is where defensible analysis begins.
Load the dataset
We will use the Iris dataset from scikit-learn, convert it to a pandas DataFrame, and standardize the column names.
Code
import pandas as pd
from sklearn.datasets import load_iris
iris = load_iris()
df = pd.DataFrame(iris.data, columns=iris.feature_names)
df["species"] = pd.Categorical.from_codes(iris.target, iris.target_names)
df.columns = ["sepal_length", "sepal_width", "petal_length", "petal_width", "species"]
df.head()
| 0 |
5.1 |
3.5 |
1.4 |
0.2 |
setosa |
| 1 |
4.9 |
3.0 |
1.4 |
0.2 |
setosa |
| 2 |
4.7 |
3.2 |
1.3 |
0.2 |
setosa |
| 3 |
4.6 |
3.1 |
1.5 |
0.2 |
setosa |
| 4 |
5.0 |
3.6 |
1.4 |
0.2 |
setosa |
What happened here?
iris.data contains the numeric measurementsiris.target contains the species labels encoded as integerspd.Categorical.from_codes() converts those encoded values into readable species names- the columns are renamed to
snake_case so later code stays clean and consistent
Save the dataset for future lessons
We will keep reusable datasets inside a local data/ folder.
Code
from pathlib import Path
Path("data").mkdir(exist_ok=True)
df.to_csv("data/iris.csv", index=False)
print("Saved dataset to: data/iris.csv")
Saved dataset to: data/iris.csv
Saving the dataset at this point helps later lessons stay reproducible and consistent.
Inspect the structure
Before plotting or summarizing, inspect the shape and variable types.
Code
print("Columns:")
print(df.columns.tolist())
print("\nShape (rows, columns):")
print(df.shape)
print("\nData types:")
print(df.dtypes)
Columns:
['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
Shape (rows, columns):
(150, 5)
Data types:
sepal_length float64
sepal_width float64
petal_length float64
petal_width float64
species category
dtype: object
Interpretation
At this stage, you should notice:
- the dataset has 150 rows
- there are 4 numeric measurement columns
species is a categorical grouping variable
That already tells us this is a small, tidy dataset that is well suited for learning exploratory analysis.
Preview the data more carefully
df.sample(8, random_state=42)
| 73 |
6.1 |
2.8 |
4.7 |
1.2 |
versicolor |
| 18 |
5.7 |
3.8 |
1.7 |
0.3 |
setosa |
| 118 |
7.7 |
2.6 |
6.9 |
2.3 |
virginica |
| 78 |
6.0 |
2.9 |
4.5 |
1.5 |
versicolor |
| 76 |
6.8 |
2.8 |
4.8 |
1.4 |
versicolor |
| 31 |
5.4 |
3.4 |
1.5 |
0.4 |
setosa |
| 64 |
5.6 |
2.9 |
3.6 |
1.3 |
versicolor |
| 141 |
6.9 |
3.1 |
5.1 |
2.3 |
virginica |
Looking at a few random rows is often more informative than only checking the first five rows.
It helps confirm that values look realistic across the dataset instead of only at the top.
Summary statistics
Code
| count |
150.000000 |
150.000000 |
150.000000 |
150.000000 |
| mean |
5.843333 |
3.057333 |
3.758000 |
1.199333 |
| std |
0.828066 |
0.435866 |
1.765298 |
0.762238 |
| min |
4.300000 |
2.000000 |
1.000000 |
0.100000 |
| 25% |
5.100000 |
2.800000 |
1.600000 |
0.300000 |
| 50% |
5.800000 |
3.000000 |
4.350000 |
1.300000 |
| 75% |
6.400000 |
3.300000 |
5.100000 |
1.800000 |
| max |
7.900000 |
4.400000 |
6.900000 |
2.500000 |
Interpretation
These summary statistics help answer early questions:
- What is the average size of each measurement?
- Which variables have wider spread?
- Are minimum and maximum values plausible?
- Do some variables appear more variable than others?
For Iris, petal measurements often show stronger separation across species than sepal measurements. We will confirm that visually next.
Species counts
Before comparing groups, verify that each group is represented.
Code
df["species"].value_counts().sort_index()
species
setosa 50
versicolor 50
virginica 50
Name: count, dtype: int64
Interpretation
This dataset is balanced across species, which makes visual comparison easier.
Exploratory visualizations
We will use modern plotting with seaborn and matplotlib, without CDI theme dependencies.
These plots are not yet final presentation graphics. Their purpose is to help us inspect the data and notice patterns worth explaining.
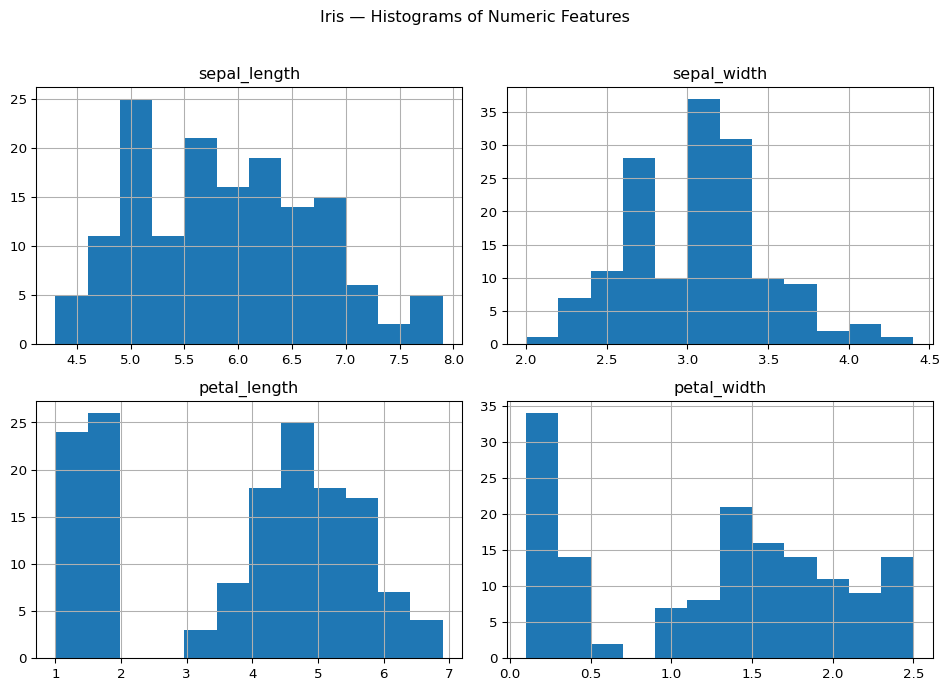
Distribution of numeric features
Code
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_theme(style="whitegrid", context="notebook")
df_long = df.melt(
value_vars=["sepal_length", "sepal_width", "petal_length", "petal_width"],
var_name="feature",
value_name="value"
)
g = sns.displot(
data=df_long,
x="value",
col="feature",
col_wrap=2,
bins=12,
height=3.6,
aspect=1.15,
facet_kws={"sharex": False, "sharey": False}
)
g.set_titles("{col_name}")
g.set_axis_labels("Value", "Count")
g.fig.suptitle("Iris — Distribution of Numeric Features", y=1.03)
plt.show()
Interpretation
These histograms help us see:
- the overall spread of each feature
- whether values are concentrated or dispersed
- whether some variables may contain overlapping or separated groups
A single histogram does not separate species, but it gives a first impression of the measurement ranges.
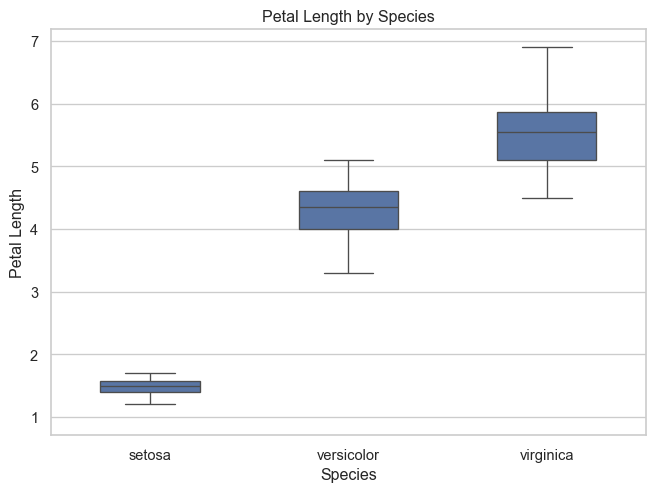
Boxplot by species
A boxplot gives a compact summary of group differences using the median, quartiles, and overall spread.
Code
fig, ax = plt.subplots(figsize=(8, 5.5))
sns.boxplot(
data=df,
x="species",
y="petal_length",
width=0.5,
fliersize=0,
ax=ax
)
ax.set_title("Petal Length by Species")
ax.set_xlabel("Species")
ax.set_ylabel("Petal Length")
plt.show()
Interpretation
This plot makes it easy to compare:
- the typical petal length in each species
- the spread within each group
- whether groups overlap
However, a boxplot hides the individual observations.
That means we cannot directly see how densely values are clustered or how points are distributed within each group.
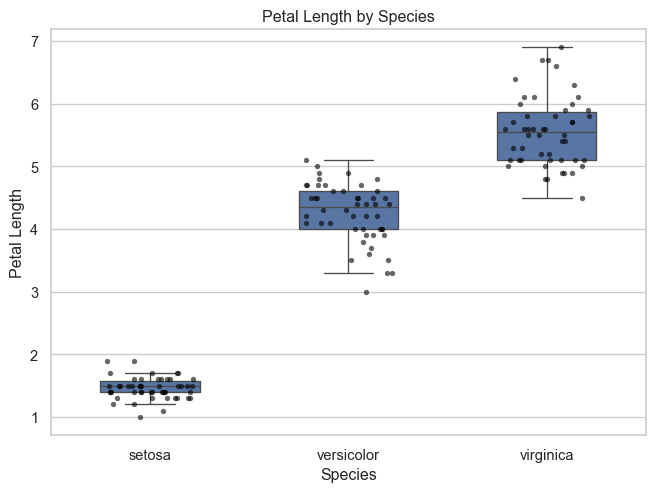
Boxplot with observed points
To make the distribution more visible, we can overlay the individual observations.
Code
fig, ax = plt.subplots(figsize=(8, 5.5))
sns.boxplot(
data=df,
x="species",
y="petal_length",
width=0.5,
fliersize=0,
ax=ax
)
sns.stripplot(
data=df,
x="species",
y="petal_length",
color="black",
alpha=0.6,
size=4,
jitter=0.22,
ax=ax
)
ax.set_title("Petal Length by Species")
ax.set_xlabel("Species")
ax.set_ylabel("Petal Length")
plt.show()
Interpretation
Adding the observed points makes the plot more informative.
Now we can see:
- how values are distributed within each species
- whether observations are tightly clustered or more dispersed
- where overlap is limited or more substantial
This provides a fuller view than the boxplot alone.
Petal length differs strongly by species.
This makes it a strong candidate for distinguishing between groups in later analysis.
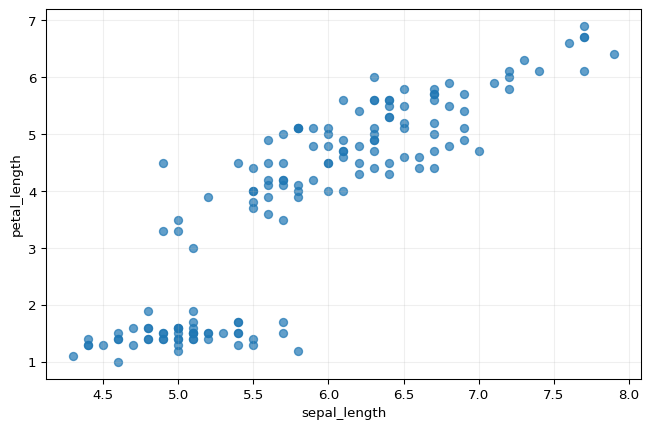
Scatter plot: sepal length vs petal length
Code
fig, ax = plt.subplots(figsize=(8, 5.5))
sns.scatterplot(
data=df,
x="sepal_length",
y="petal_length",
hue="species",
s=70,
alpha=0.8,
ax=ax
)
ax.set_title("Sepal Length vs Petal Length")
ax.set_xlabel("Sepal Length")
ax.set_ylabel("Petal Length")
plt.show()
Interpretation
Scatter plots help us inspect relationships between two variables.
Here, the species begin to separate into visible clusters, especially when petal length is involved.
This is a strong exploratory signal that some measurements carry more discriminatory information than others.
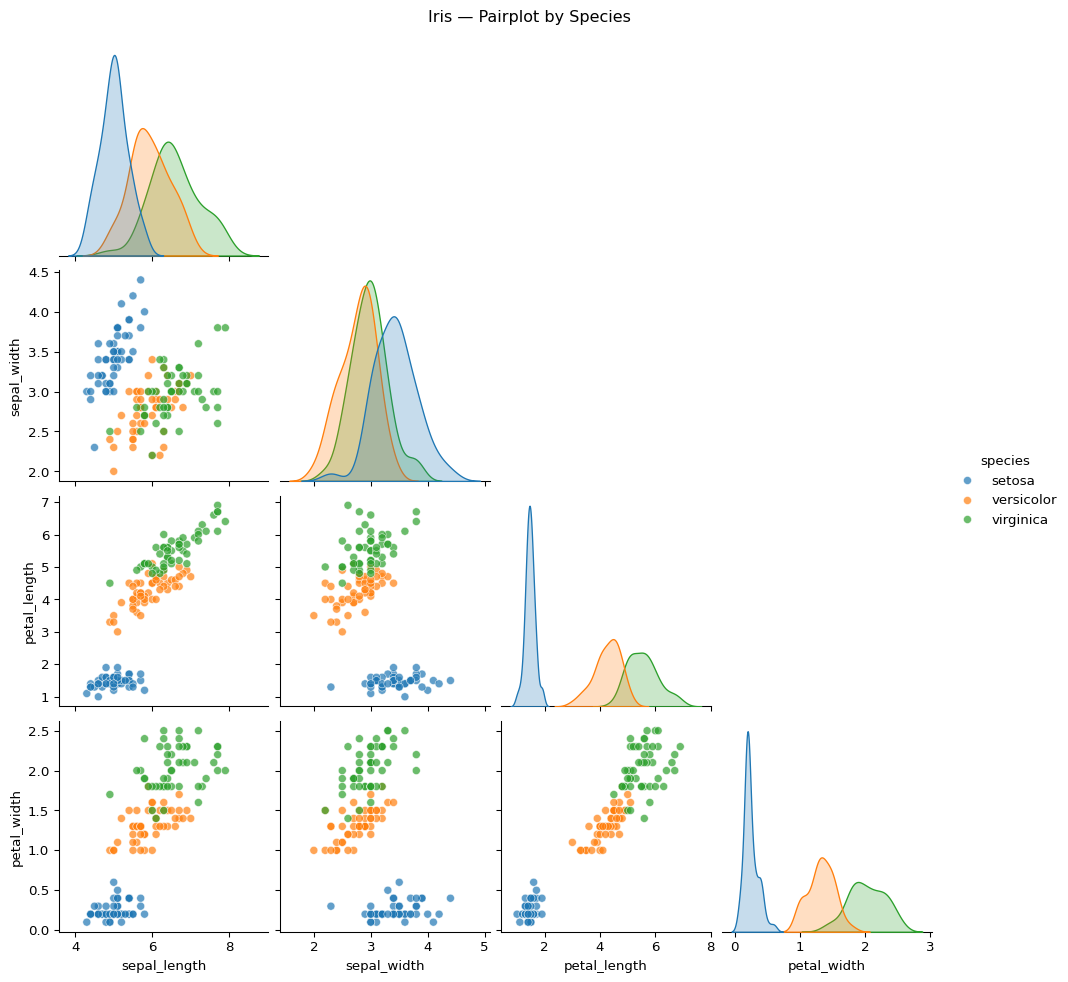
Pairwise relationships
Code
g = sns.pairplot(
df,
hue="species",
corner=True,
diag_kind="hist",
plot_kws={"alpha": 0.7, "s": 45}
)
g.fig.suptitle("Iris — Pairwise Relationships by Species", y=1.02)
plt.show()
Interpretation
The pairplot provides a broader view of the dataset:
- which variable pairs show clear separation
- where clusters overlap
- which variables appear more useful for distinguishing species
Create a simple derived feature
Feature creation is often part of exploration.
Here, we define a new variable, petal_area, as a simple combination of petal length and petal width.
Code
df["petal_area"] = df["petal_length"] * df["petal_width"]
df.head()
| 0 |
5.1 |
3.5 |
1.4 |
0.2 |
setosa |
0.28 |
| 1 |
4.9 |
3.0 |
1.4 |
0.2 |
setosa |
0.28 |
| 2 |
4.7 |
3.2 |
1.3 |
0.2 |
setosa |
0.26 |
| 3 |
4.6 |
3.1 |
1.5 |
0.2 |
setosa |
0.30 |
| 4 |
5.0 |
3.6 |
1.4 |
0.2 |
setosa |
0.28 |
This does not mean the new feature is automatically better.
It illustrates how exploratory analysis can lead to new candidate variables.
Exercise
Try the following:
- Print the last five rows of the dataset.
- Compute the mean of
petal_area for each species.
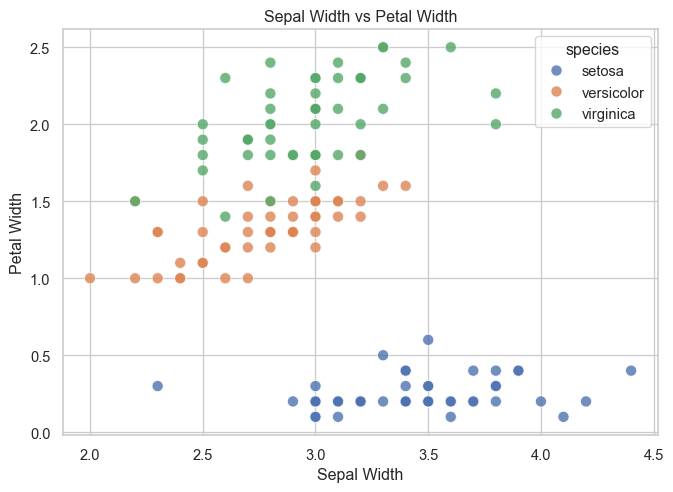
- Create a scatter plot of
sepal_width versus petal_width.
- Write one sentence describing which feature seems most useful for separating species.
Code
print("Last 5 rows:")
print(df.tail())
print("\nMean petal_area by species:")
print(df.groupby("species", observed=False)["petal_area"].mean())
fig, ax = plt.subplots(figsize=(8, 5.5))
sns.scatterplot(
data=df,
x="sepal_width",
y="petal_width",
hue="species",
s=70,
alpha=0.8,
ax=ax
)
ax.set_title("Sepal Width vs Petal Width")
ax.set_xlabel("Sepal Width")
ax.set_ylabel("Petal Width")
plt.show()
Last 5 rows:
sepal_length sepal_width petal_length petal_width species \
145 6.7 3.0 5.2 2.3 virginica
146 6.3 2.5 5.0 1.9 virginica
147 6.5 3.0 5.2 2.0 virginica
148 6.2 3.4 5.4 2.3 virginica
149 5.9 3.0 5.1 1.8 virginica
petal_area
145 11.96
146 9.50
147 10.40
148 12.42
149 9.18
Mean petal_area by species:
species
setosa 0.3656
versicolor 5.7204
virginica 11.2962
Name: petal_area, dtype: float64
Interpretation:
Petal-based features provide clearer separation between species than sepal-based features.
CDI Insight
Exploration is not just about making plots.
It is about understanding how the dataset behaves before it is used to support conclusions.
A responsible analyst does not move directly from data loading to modeling.
They pause, inspect, compare, and question — and only then move forward.
That habit is a foundation of reliable analysis.
Summary
In this lesson, you:
- loaded the Iris dataset into a pandas DataFrame
- standardized column names
- saved the dataset to
data/iris.csv
- inspected structure, data types, and summary statistics
- used exploratory visualizations to examine distributions and group differences
- created a simple derived feature for further exploration
Next Step
Next, we clean and prepare the dataset for analysis.
Fisher, Ronald A. 1936. “The Use of Multiple Measurements in Taxonomic Problems.” Annals of Eugenics.